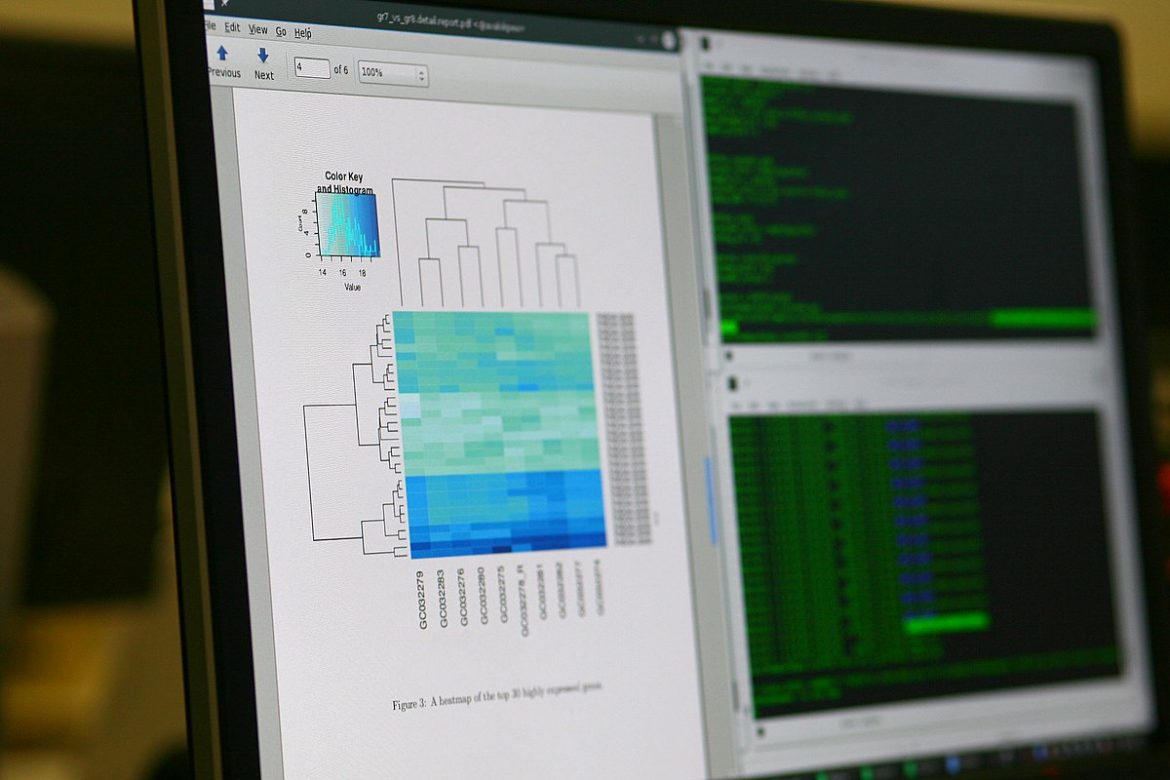
Human full genome data analysis at the Flemish Super Computer Center
The analysis of human full genome data is a computationally demanding task. Typically, raw data amounts to 80GB of compressed short reads per sample, which first has to be mapped to a reference genome and then analyzed for variant detection. Even on a performance cluster as the one currently available at the UZ Leuven, the whole process may take several days, or even weeks. To cope with this computational burden, in collaboration with the Flemish Super Computer Center (VSC), the Genomics Core is now performing human full-genome variant calling analysis on the high performance computing infrastructure of the VSC. The results for the first full-genome analyses are encouraging, and the whole pipeline execution is now measured in hours rather than days.